MIACYC


The MIACYC PROJECT
Plants are an inexhaustible source of pharmaceutical compounds, essential component of our pharmacopeia. Unfortunately, these natural products (NP) accumulate in minute amounts in plants while their complex structures render total chemical syntheses highly impractical, leading to the overexploitation of natural resources.
Developing new and complementary approaches to produce highly valuable NPs is thus mandatory to secure supply at midterms while preserving natural resources. This can be notably envisaged through the conversion of abundant and simpler NP precursors that can be extracted from plants commonly grown in Europe.
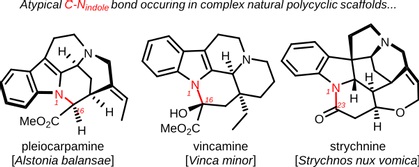
It’s particularly true for the monoterpenoid indole alkaloids (MIAs), a class of NP used mostly in chemotherapies which are currently produced by semisynthesis resulting in limited supplies at exhorbitant market prices. MIACYC will focus on the elucidation of an atypical C-N indole cyclization of MIAs found in the biosynthesis of three emblematic MIAs : pleiocarpamine, vincamine and strychnine. These natural products endow with pharmacological properties and occur in three plant models (see below).
While the enzymatic reactions catalyzing the C-Nindole cyclization are still poorly documented, MIACYC aims identifying the corresponding biosynthetic reactions allowing the development of a new pipeline of pathway elucidation and opening doors for a sustainable production of these compounds and their potential derivatives through yeast metabolic engineering.


Our plant models

producing vincamine with nootropic effect

producing pleiocarpamine, an anticholinergic

producing strychnine, a stimulant
Our objectives
Identification of new scaffolds of MIA at the cellular level using metabolomic innovative tools
Identification of the key enzymes involved in the biosynthesis of C-Nindole MIAs
Production in microbes of targeted MIA using genes encoding enzymes involved in C-Nindole alkaloid metabolism
